For a more complete and up to date list, see Ryan's Google Scholar Profile
2026
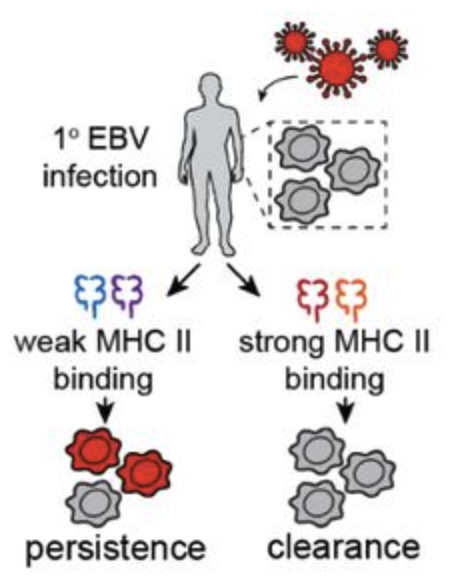
[53] Population-scale sequencing resolves correlates and determinants of latent Epstein-Barr Virus infection
S Nyeo, EM Cumming, OS Burren, MS Pagadala, JC Gutierrez, TA Ali, LC Kida, Y Chen, F Hu, B Hollis, M Fabre, S MacArthur, Q Wang, LS Ludwig, KK Dey, S Petrovski*, RS Dhindsa*, CA Lareau*.
Nature. (2025).
[52] Phenome-wide analysis of copy number variants in 470,727 UK Biobank genomes
XZ Zou, F Hu, H Lou, OS Burren, X Li, K Megy, E Wheeler, Q Wu, SS Atanur, M Karpinski, D Loesch, Z Fairhurst-Hunter, SVV Deevi, E Oerton, S Wen, X Jiang, C Salvoro, J Mitchell, A Nag, B Hollis, A O’Neill, AstraZeneca Genomics Initiative, J Harrow, S MacArthur, S Wasilewski, S O’Dell, L Tian, KR Smith, G del Angel, M Fabre, RS Dhindsa, Q Wang, S Petrovski, K Carss
Nature. (2026).
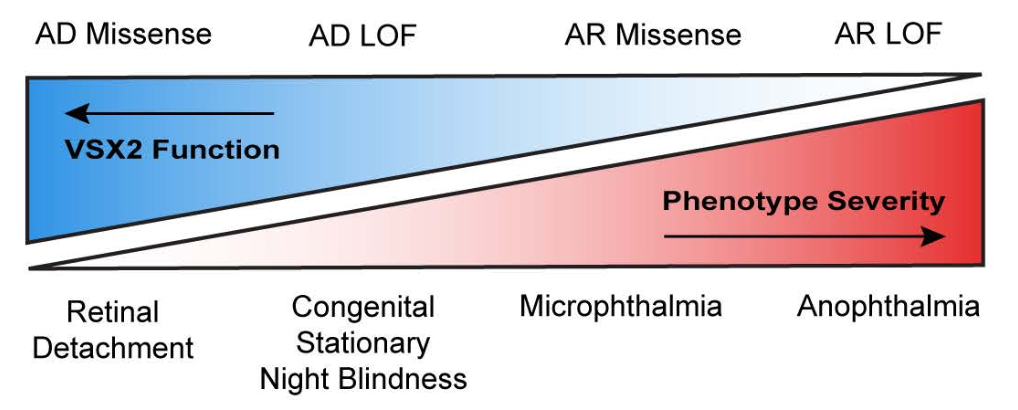
[51] Rare heterozygous missense variants in VSX2 are associated with retinal detachment
DC Brock*, JS Dhindsa*, Y Chen, V Ravanmehr, J Mitchell, F Hu, X Li, L Nandigam, Q Wang, K Wu, JC Butts, HS Dhindsa, BJ Frankfort, NM Tran, S Petrovski, RS Dhindsa
PLoS Genetics. (2026).
2025
[50] Haploinsufficiency of ITSN1 is associated with a substantial increased risk of Parkinson’s disease
TP Spargo, CF Sands, IR Juan, J Mitchell, V Ravanmehr, JC Butts, RB De-Paula, Y Kim, F Hu, Q Wang, D Vitsios, M Garg, M Messa, G del Angel, DG Calame, H Saade, L Robak, B Hollis, HY Zoghbi, J Shulman, S Petrovski, I Al-Ramahi, I Tachmazidou, RS Dhindsa.
Cell Reports. (2025).

[49] Whole-genome sequencing of 490,640 UK Biobank participants
The UK Biobank Whole-Genome Sequencing Consortium
Nature. (2025).
[48] Minding the synapse: A multi-omic approach reveals hidden regulators of neurodevelopment
A Ljungdahl, RS Dhindsa.
Cell Systems. (2025).
[47] Characterising the contribution of rare protein-coding germline variants to prostate cancer risk and severity in 37,184 cases
J Mitchell, N Camacho, P Shea, H Stopsack, V Joseph, O Burren, RS Dhindsa, Abhishek Nag, et al.
Nature Communications. (2025).
[46] Genome-wide prediction of dominant and recessive neurodevelopmental disorder risk genes
RS Dhindsa, B Weido, JS Dhindsa, AJ Shetty, C Sands, S Petrovski, D Vitsios, AW Zoghbi.
American Journal of Human Genetics. (2025)

[45] Comparative analysis of the Mexico City Prospective Study and the UK Biobank identifies ancestry-specific effects on clonal hematopoiesis
S Wen, P Kuri-Morales, F Hu, A Nag, I Tachmazidou, S Deevi, H Taiy, K Smith, DP Loesch, OS Burren, RS Dhindsa, S Wasilewski, et al.
Nature Genetics. (2025).
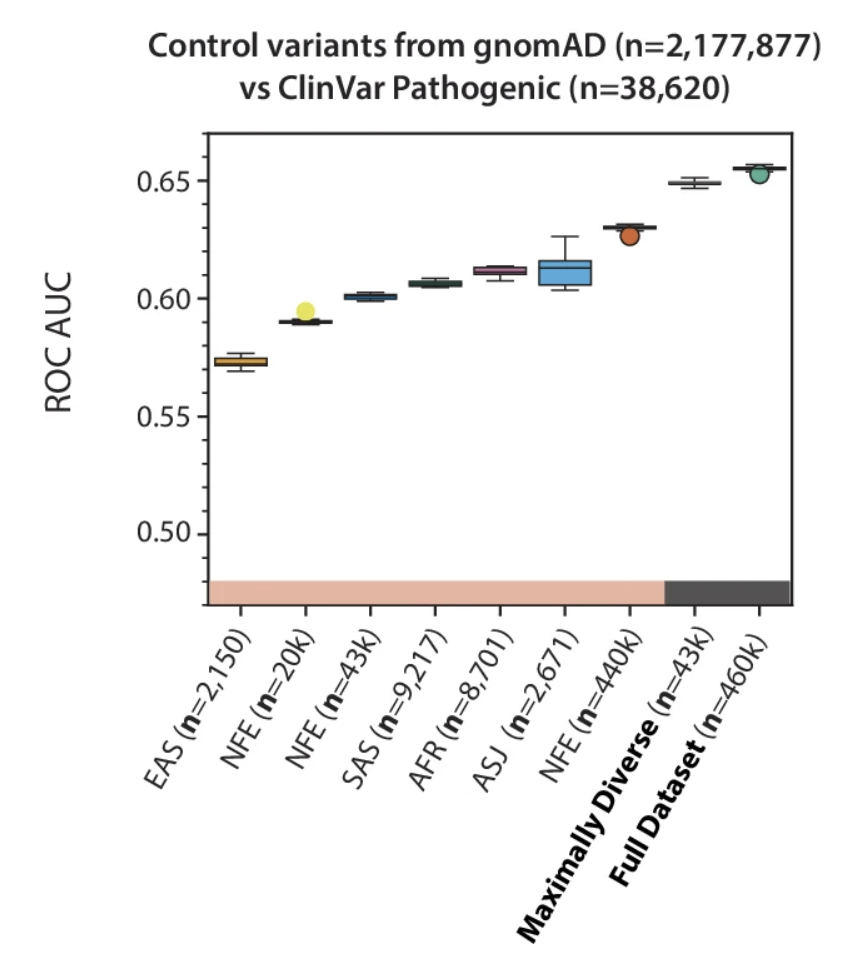
[44] Diverse ancestral representation improves genetic intolerance metrics
AL Han, CF Sands, D Matelska, JC Butts, V Ravanmehr, F Hu, EV Gonzalez, N Katsanis, CD Bustamante, Q Wang, S Petrovski, D Vitsios, RS Dhindsa.
Nature Communications. (2025).
2024
[42] A single-cell transcriptomic map of the developing Atoh1 lineage identifies neural fate decisions and neuronal diversity in the hindbrain
JC Butts, SWu, MA Durham, RS Dhindsa, J Revelli, MC Ljungberg, O Saulnier, ME McLaren, MD Taylor, HY Zoghbi.
Developmental Cell. (2024).
[41] Disease prediction with multi-omics and biomarkers empowers case–control genetic discoveries in the UK Biobank
M Garg, M Karpinski, D Matelska, L Middleton, J Mitchell, A O'Neill, Q Wang, AR Harper, RS Dhindsa, S Petrovski, D Vitsios.
Nature Genetics. (2024).
[40] Phenome-wide identification of therapeutic genetic targets, leveraging knowledge graphs, graph neural networks, and UK Biobank data
L Middleton, I Melas, C Vasavda, A Raies, B Rozemberczki, RS Dhindsa, JS Dhindsa, B Weido, Q Wang, AR Harper, G Edwards, S Petrovski, D Vitsios.
Science Advances. (2024).
[39] Genetic architecture of telomere length in 462,666 UK Biobank whole-genome sequences
OS Burren*, RS Dhindsa*, S Deevi*, S Wen, A Nag, J Mitchell, F Hu, KR Smith, N Razdan, H Olsson, A Platt, D Vitsios, Q Wu, AstraZeneca Genomics Initiative, V Codd, CP Nelson, NJ Samani, RE March, S Wasilewski, K Carss, M Fabre, Q Wang, MN Pangalos, S Petrovski.
Nature Genetics. (2023).
[38] An expanded genomic database for identifying disease-related variants
RS Dhindsa, S Petrovski.
Nature. (2024).
2023
[37] Rare variant associations with plasma protein levels in the UK Biobank
RS Dhindsa*, OS Burren*, BB Sun*, BP Prins, D Matelska, E Wheeler, J Mitchell, E Oerton, VA Hristova, KR Smith, K Carss, S Wasilewski, AR Harper, DS Paul, MA Fabre, H Runz, C Viollet, B Challis, A Platt, AstraZeneca Genomics Initiative, D Vitsios, EA Ashley, CD Whelan, MN Pangalos, Q Wang, S Petrovski.
Nature. (2023)

[36] Plasma proteomic associations with genetics and health in the UK Biobank
B Sun, ... AstraZeneca Genomics Initiative, ..., JD Szustakowski, BW Gibson, MR Miller, CD Whelan.
Nature. (2023).
[35] Neurodevelopmental deficits and cell-type-specific transcriptomic perturbations in a mouse model of HNRNPU haploinsufficiency
SA Dugger*, RS Dhindsa*, G Sampaio, E Rafikian, S Petri, J Teoh, J Ye, S Colombo, M Yang, M Boland, W Frankel, DB Goldstein.
PLoS Genetics. (2023).

[34] Literature-based predictions of Mendelian disease therapies
CA Deisseroth, W Lee, J Kim, H Jeong, RS Dhindsa, J Wang, HY Zoghbi, Z Li
American Journal of Human Genetics. (2023).
[33] Atoh1 drives the heterogeneity of the pontine nuclei neurons and promotes their differentiation
S Wu, JC Butts, MS Caudill, JP Revelli, RS Dhindsa, MA Durham, HY Zoghbi.
Science Advances. (2023).
[32] Codon affinity in mitochondrial DNA shapes evolutionary and somatic fitness
CA Lareau, Y Yin, JC Gutierrez, RS Dhindsa, A Gribling-Burrer, Y Hsieh, L Nitsch, FA Buquicchio, T Abay, S Zielinski, RR Stickels, JC Ulirsch, P Yan, F Wang, Z Miao, K Sandor, B Daniel, V Liu, Q Wang, F Hu, KR Smith, S Deevi, P Maschmeyer, S Petrovski, RP Smyth, WJ Greenleaf, A Kundaje, M Munschauer, S Ludwig, A Satpathy.
bioRxiv. (2023).
[31] Epilepsy in a mouse model of GNB1 Encephalopathy arises from altered potassium channel (GIRK) signaling and is alleviated by a GIRK inhibitor
S Colombo, HP Reddy, S Petri, DJ Williams, B Shalomov, RS Dhindsa, S Gelfman, D Krizay, AK Bera, M Yang, Y Peng, C Makinson, MJ Boland, WN Frankel, DB Goldstein, N Dascal.
Frontiers in Cellular Neuroscience. (2023).
[30] Studying ultra-rare variants in STX1A uncovers a novel neurodevelopmental disorder
EV Gonzalez and RS Dhindsa.
European Journal of Human Genetics. (2023).
[29] Effects of protein-coding variants on blood metabolite measurements and clinical biomarkers in the UK Biobank
A Nag, RS Dhindsa, L Middleton, Xiao Jiang, D Vitsios, E Wigmore, EL Allman, A Reznichenko, K Carss, KR Smith, Q Wang, B Challis, DS Paul, AR Harper, S Petrovski.
American Journal of Human Genetics. (2023).
[28] Genome-wide identification and phenotypic characterization of seizure-associated copy number variations in 741,075 individuals
L Montanucci, D Lewis-Smith, RL Collins, ..., Epi25 Collaborative, I Helbig, C Leu, D Lal.
Nature Communications. (2023).
[27] Epilepsies of presumed genetic etiology show enrichment of rare variants that occur in the general population
L Bundalian, YY Su, S Chen, A Velluva, ..., Epi25 Collaborative.
American Journal of Human Genetics. (2023).
2022
[26] A minimal role for synonymous variation in human disease
RS Dhindsa, Q Wang, D Vitsios, OS Burren, F Hu, JE DiCarlo, L Kruglyak, DG MacArthur, ME Hurles, S Petrovski.
American Journal of Human Genetics. (2022).
[25] DrugnomeAI is an ensemble machine-learning framework for predicting druggability of candidate drug targets
A Raies, E Tulodziecka, J Stainer, L Middleton, RS Dhindsa, P Hill, O Engkvist, AR Harper, S Petrovski, D Vitsios.
Communications Biology. (2022)
[24] Human genetic evidence supports MAP3K15 as an obesity-independent therapeutic target for diabetes
A Nag*, RS Dhindsa*, J Mitchell, C Vasavda, AR Harper, et al.
Science Advances. (2022).
[23] Identification of the NRF2 transcriptional network as a therapeutic target for trigeminal neuropathic pain
C Vasavda, R Xu, J Liew, R Kothari, RS Dhindsa, E Semenza, B Paul, et al.
Science Advances. (2022).
[22] Cancer-driving mutations are enriched in genic regions intolerant to germline variation
D Vitsios*, RS Dhindsa*, J Mitchell, D Matelska, Z Zou, J Armenia, Q Wang, B Sidders, AR Harper, S Petrovski.
Science Advances. (2022).
[21] Biliverdin reductase bridges focal adhesion kinase to Src to modulate synaptic signaling
C Vasavda, ER Semenza, J Liew, R Kothari, RS Dhindsa, S Shanmukha, A Lin, R Tokhunts, C Ricco, AM Snowman, LK Albacarys, F Pastore, C Ripoli, C Grassi, E Barone, MD Kornberg, X Dong, BD Paul, SH Snyder.
Science Signaling. (2022).
[20] Ancestry adjustment improves genome-wide estimates of regional intolerance
T Hayeck, N Stong, E Baugh, RS Dhindsa, TN Turner, A Malakar, Y Duan, I Ionita-Laza, DB Goldstein, AS Allen.
Genetics. (2022).
[19] Association of ultra-rare coding variants with genetic generalized epilepsy: A case–control whole exome sequencing study
M Koko, JE Motelow, KE Stanley, DR Bobbili, RS Dhindsa, P May, Canadian Epilepsy Network, Epi4K Consortium, Epilepsy Phenome/Genome Project, EpiPGX Consortium, EuroEPINOMICS-CoGIE Consortium.
Epilepsia. (2022).
2021
[18] High-impact rare genetic variants in severe schizophrenia
AW Zoghbi, RS Dhindsa, TE Goldberg, A Mehralizade, JE Motelow,X Wang, A Alkelai, MB Harms, JA Lieberman, S Markx. D Goldstein.
PNAS. (2021).
[17] Rare variant contribution to human disease in 281,104 UK Biobank exomes
Q Wang*, RS Dhindsa*, K. Carss*, AR Harper, A Nag, I Tachmazidou, D Vitsios, SVV Deevi, A Mackay, D Muthas, Michael Hühn, S Monkley, H Olsson, S Wasilewski, KR Smith, R March, A Platt, C Haefliger, S Petrovski.
Nature. (2021).

[16] Distinct gene-set burden patterns underlie common generalized and focal epilepsies
Epi25 Consortium.
EBioMedicine, (2021).
[15] Sub-genic intolerance, ClinVar, and the epilepsies: A whole-exome sequencing study of 29,165 individuals
Epi25 Consortium.
American Journal of Human Genetics. (2021).
[14] Identification of a missense variant in SPDL1 associated with idiopathic pulmonary fibrosis
RS Dhindsa, J Mattsson, A Nag, Q Wang, LV Wain, R Allen, EM Wigmore, K Ibanez, D Vitsios, SV Deevi, S Wasilewski, M Karlsson,G Lassi, H Olsson, D Muthas, S Monkley, A Mackay, L Murray, S Young, C Haefliger, FinnGen Consortium, TM Maher, MG Belvisi, G Jenkins, P Molyneaux, A Platt, S Petrovski.
Communications Biology. (2021).

[13] Prioritizing non-coding regions based on human genomic constraint and sequence context with deep learning
D Vitsios, RS Dhindsa, L Middleton, AB Gussow, S Petrovski.
Nature Communications. (2021).
[12] A Transcriptome‐Based Drug Discovery Paradigm for Neurodevelopmental Disorders
RS Dhindsa, AW Zoghbi, DK Krizay, C Vasavda, DB Goldstein.
Annals of Neurology. (2021).

2020
[11] TMPRSS2 Transcriptional Inhibition as a Therapeutic Strategy for COVID-19
X Wang, RS Dhindsa, G Povysil, A Zoghbi, J Motelow, J Hostyk, N Nickols, M Rettig, DB Goldstein.
Preprints. (2020).
[10] Natural selection shapes codon usage in the human genome
RS Dhindsa, BR Copeland, AM Mustoe, DB Goldstein.
American Journal of Human Genetics. (2020).

Pre-2020
[9] Ultra-rare genetic variation in the epilepsies: a whole-exome sequencing study of 17,606 individuals
Epi25 consortium.
American Journal of Human Genetics. (2019).
[8] meaRtools: An R package for the analysis of neuronal networks recorded on microelectrode arrays
S Gelfman, Q Wang, Y Lu, D Hall, CD Bostick, RS Dhindsa, M Halvorsen, KM McSweeney, E Cotterill, T Edinburgh, MA Beaumont, WN Frankel, S Petrovski, AS Allen, MJ Boland, DB Goldstein, SJ Eglen.
PLoS computational biology. (2018).
[7] Orion: detecting regions of the human non-coding genome that are intolerant to variation using population genetics
AB Gussow, BR Copeland, RS Dhindsa, Q Wang, S Petrovski, WH Majoros, AS Allen, DB Goldstein.
PloS one. (2017).
[6] Molecular Architecture and Neurobiology of the Epilepsies
RS Dhindsa, DH Lowenstein, DB Goldstein.
Genomics, Circuits, and Pathways in Clinical Neuropsychiatry. (2016).
[5] Schizophrenia: From genetics to physiology at last
RS Dhindsa, DB Goldstein.
Nature. (2016).

[4] Genetic Discoveries Drive Molecular Analyses and Targeted Therapeutic Options in the Epilepsies
RS Dhindsa, DB Goldstein.
Current neurology and neuroscience reports. (2015).
[3] Exome sequencing results in successful riboflavin treatment of a rapidly progressive neurological condition
S Petrovski, V Shashi, S Petrou, K Schoch, KM McSweeney, RS Dhindsa, B Krueger, R Crimian, LE Case, R Khalid, MA El-Dairi, Y Jiang, MA Mikati, DB Goldstein.
Molecular Case Studies. (2015).
[2] Whole-exome sequencing in undiagnosed genetic diseases: interpreting 119 trios
X Zhu, S Petrovski, P Xie, EK Ruzzo, Y Lu, KM McSweeney, B Ben-Zeev, A Nissenkorn, Y Anikster, D Oz-Levi, RS Dhindsa, Y Hitomi, K Schoch, RC Spillmann, G Heimer, D Marek-Yagel, M Tzadok, Y Han, G Worley, J Goldstein, Y Jiang, D Lancet, E Pras, V Shashi, D McHale, AC Need, DB Goldstein.
Genetics in Medicine. (2015).
[1] Epileptic encephalopathy-causing mutations in DNM1 impair synaptic vesicle endocytosis.
RS Dhindsa, SS Bradrick, X Yao, EL Heinzen, S Petrovski, BJ Krueger, MR Johnson, WN Frankel, S Petrou, RM Boumil, DB Goldstein.
Neurology Genetics. (2015).
